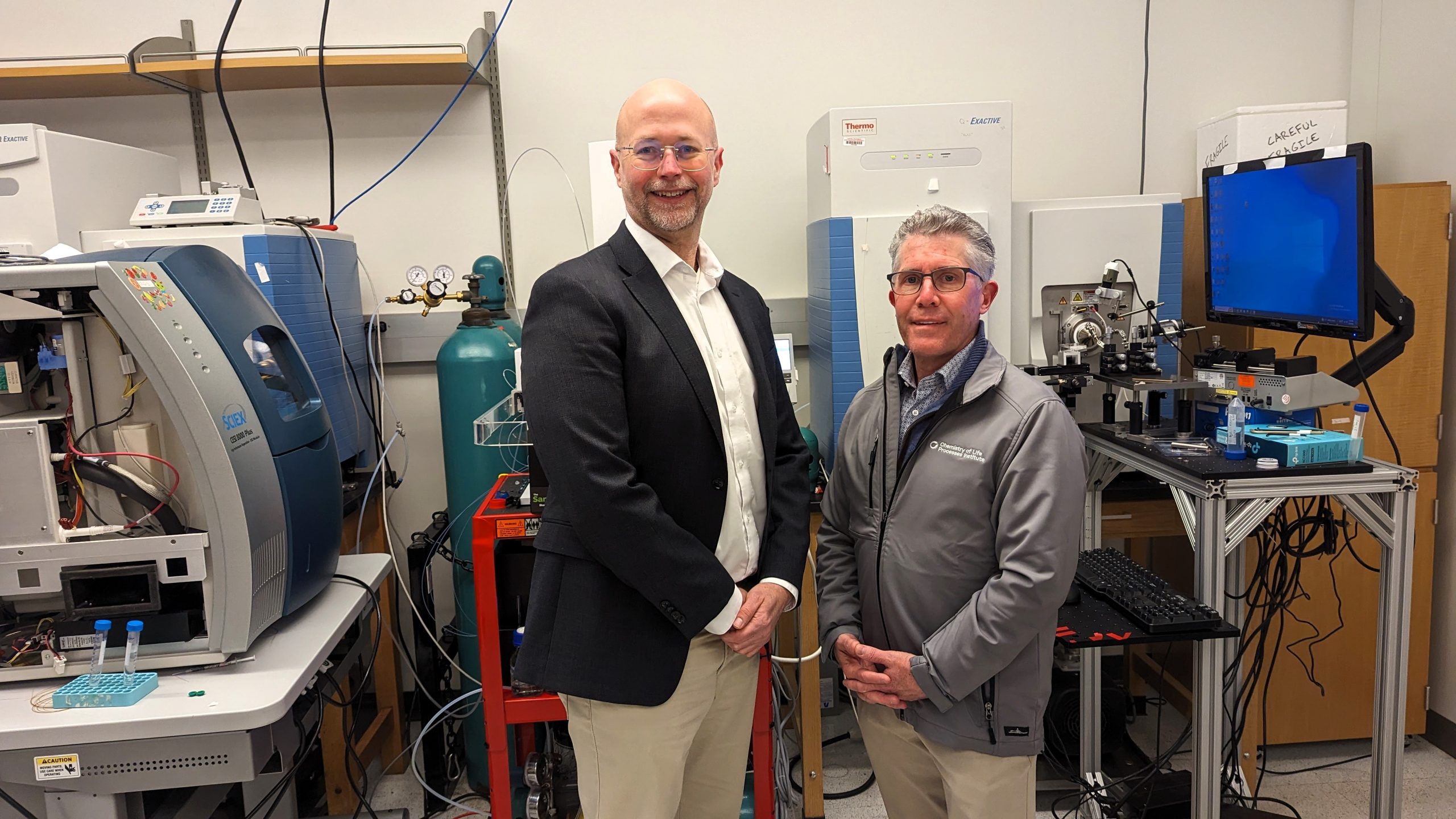
As is the case with many other diseases, a protein is thought to be a key driver of Parkinson’s disease (PD). Thanks to a two-year $700,000 grant from The Michael J. Fox Foundation for Parkinson’s Research (MJFF), researchers in Northwestern’s Chemistry […]
Neil Kelleher, Walter and Mary Elizabeth Glass Professor of Chemistry, Molecular Biosciences, and Medicine at Northwestern University, will present his groundbreaking work and insights at US HUPO 2024 in Portland, Oregon next week as the 2024 recipient of the Donald […]
By Olivia Dimmer Nov 9, 2023 Investigators led by Neil Kelleher, PhD, professor of Medicine in the Division of Hematology and Oncology and of Biochemistry and Molecular Genetics, have developed an automated technique for imaging and identifying proteoforms in ovarian cancer tissue, according to results published in Nature Communications. […]
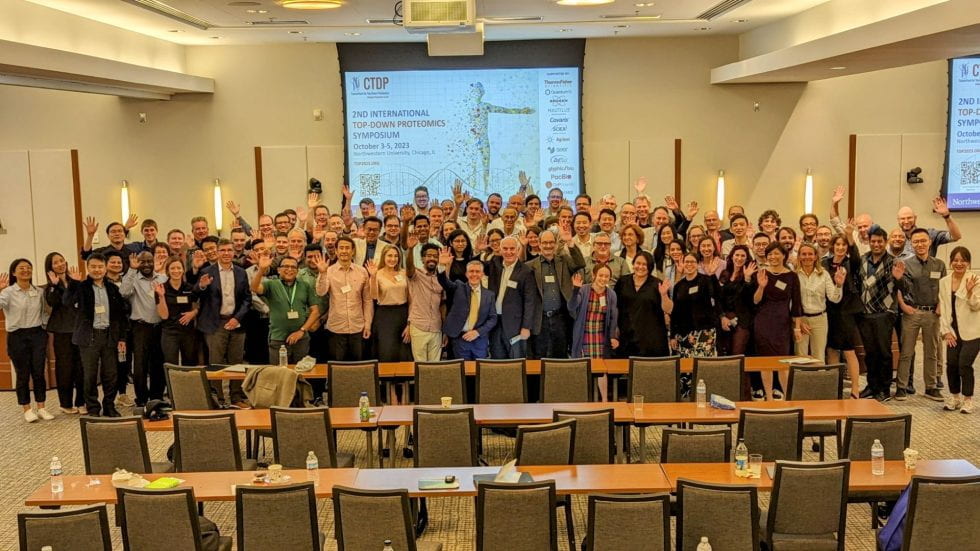
One hundred and sixty world leaders in proteomics, proteoform biology, cell biology and genomics gathered October 3-5, 2023, at Northwestern’s Prentice Women’s Hospital to present their latest research findings and discuss next-generation proteomics at the second International Top-Down Proteomics […]
Utilizing advanced top-down proteomics approaches, Northwestern Proteomics researchers and collaborators in the Jewett Lab identified a proteoform responsible for increasing the speed of a well-known metabolic enzyme, Triosephosphate Isomerase (TPI) that drives energy production in the cell. While […]
What do wrestling and research have in common? “Hard work and discipline,” says student-athlete Troy Fisher, one of four Northwestern wrestling team members who interned for the Kelleher Research Group last summer. Fisher and fellow students Jon Halvorsen, Andrew Davison, […]
The Society for Analytical Chemists of Pittsburgh (SACP) and the Pittsburgh Conference on Analytical Chemistry and Applied Spectroscopy (Pittcon), will present the Pittsburgh Analytical Chemistry Award to Northwestern’s Neil L. Kelleher, PhD, the Walter and Mary E. Glass Professor of […]
Two recent papers by Northwestern Proteomics researchers demonstrate the critical importance of proteoform analysis (the study of all of the different forms of proteins in the human body) in developing and refining treatments for cancer and other diseases. […]
Northwestern University’s Chemistry of Life Processes Institute will host the second International Top-Down Proteomics Symposium which will be held in Chicago, IL, October 3-5, 2023. The Symposium, a Consortium for Top-Down Proteomics event, will gather the global community to present and discuss the latest […]
We have a new presentation explaining the needs and the value of the Human Proteoform Project. Watch Neil Kelleher take you through the project in 12 minutes. Original post can be found here on The Consortium for Top-Down […]